Computational neuroscience methods
Tools and models for analyzing neural circuits and designing visual stimuli
Geometry of color spaces (dreye)
Color vision represents a vital aspect of perception that ultimately enables a wide variety of species to thrive in the natural world. However, unified methods for constructing chromatic visual stimuli in a laboratory setting are lacking. Here, we present stimulus design methods and an accompanying programming package to efficiently probe the color space of any species in which the photoreceptor spectral sensitivities are known. Our hardware-agnostic approach incorporates photoreceptor models within the framework of the principle of univariance. This enables experimenters to identify the most effective way to combine multiple light sources to create desired distributions of light, and thus easily construct relevant stimuli for mapping the color space of an organism. We include methodology to handle uncertainty of photoreceptor spectral sensitivity as well as to optimally reconstruct hyperspectral images given recent hardware advances. Our methods support broad applications in color vision science and provide a framework for uniform stimulus designs across experimental systems.
Exploiting colour space geometry for visual stimulus design across animals, Phil. Trans. R. Soc. B, 2022
Probabilistic circuit model analysis
Sensory circuits are complex systems that process information through feedforward and recurrent interactions among a set of neurons. While these circuits have been studied extensively, modeling their underlying mechanisms and computations can be challenging when not all neurons in the circuit are recorded from and when synapse information is incomplete or inaccurate. In this study, we propose a probabilistic framework that predicts the responses of unobserved neurons using anatomical constraints. We demonstrate the effectiveness of our approach on both simulated data using randomly generated connectomes with varying complexities and real neural data from the fruit fly optic lobe. Our approach accurately predicts the responses of unobserved neurons in simulated and real data. Given our probabilistic framework, we are also able to infer what types of experiments are optimal for subsequent experiments. Overall, our method provides a useful tool for modeling the complex mechanisms underlying sensory information processing in biological circuits.
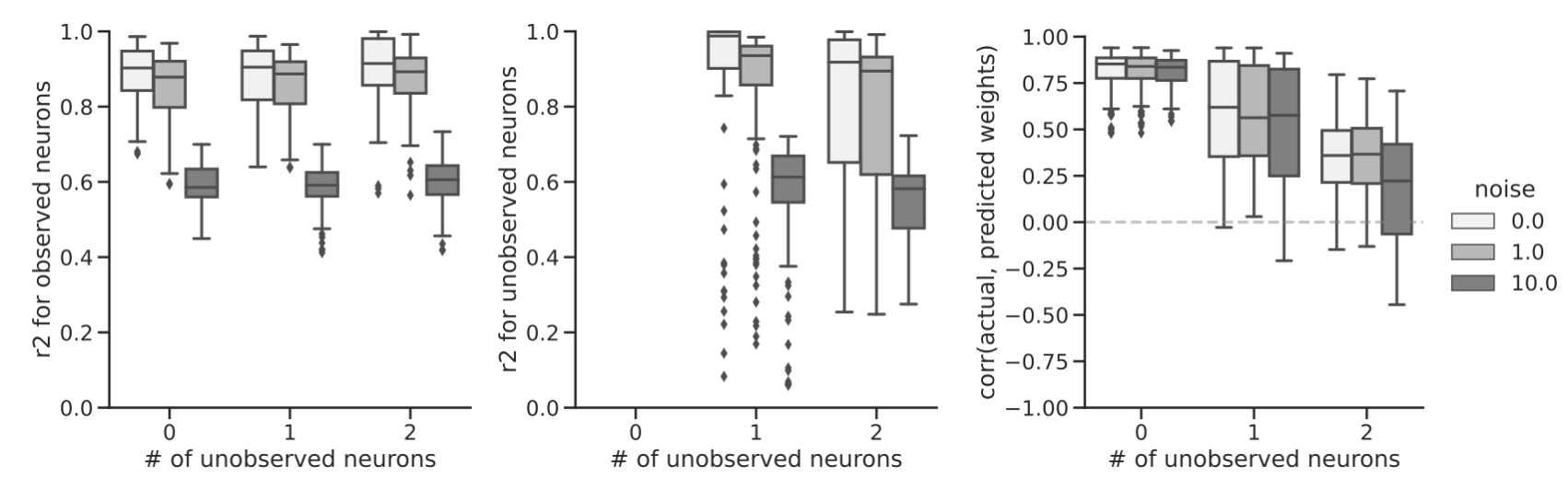
Presented at Columbia Neurotheory Meeting, 2020 and 2021
Encoding models (scidoggo)
This coding project is a collection of different models and data tools that I have created, modified, and used for my various research projects. Included in the package are various types of constraint convex optimization procedures, interpolation models, nonlinear encoding models, and probabilistic models for circuit model analysis. The project is still under development and can be found on GitHub.