
medical AI · data · neuroML
blog
Where my work moved
What changed for me with “AI” is not just speed. I write fewer first drafts by hand now, but I spend more time specifying what I want, pruning what the model produced, and checking that I still understand it. Am I being more thorough than before? I don’t quite know. My old mistakes versus the model’s are usually different. Mine tended to come from moving too fast through an idea: indexing mistakes, edge cases I did not split out, or a spec that was still half in my head (I used to have a lot more just in my head than nowadays). The model fails differently. It swallows errors, picks the wrong abstraction level, or produces far more structure than the task actually needed.
The shift became clear once I split my coding work into two broad categories: exploratory analysis and production code. In exploratory analysis, I used to poke around in notebooks until the shape of the analysis emerged. Now I usually specify the analysis up front, in marimo rather than Jupyter (the model can generate and debug that more cleanly), let it build a first pass, and then spend my time pruning, checking, and editing the notebook into something coherent. The exploratory loop is still there. I just write less of the first pass myself, and I produce (i.e. let the “AI” generate) and throw away more analysis side quests.
In production code, I used to test ideas more directly in code or in small exploratory fragments (often in notebooks). Now I start with an exploratory discussion with the “AI” and then write a specification, test cases, and finally implement. The main problem is not producing usable code anymore, but rather getting the abstractions right and making my assumptions explicit. I still want to understand the generated code well enough to “own” it later. Because code is cheap to generate, I am also more willing to throw it away and restart from a cleaner spec instead of repairing an existing implementation.
The setup tax for automating complex tasks
The model’s ability to cheaply produce first drafts made it worthwhile to automate recurring semi-manual tasks that I previously ignored.
For example, I recently implemented tooling to generate data-heavy PowerPoint slides that had to follow an internal style guide and a specific template. Building the first version of that workflow took longer than making the slides by hand. I needed tooling for the template, tooling for the data warehouse, and a configuration format that allowed the model to describe layout, figures, and text without directly interacting with PowerPoint.
The early outputs of this workflow were buggy and not pretty (e.g. bad spacing, wrong sizing, too much stuff on one slide, overlapping text). Simply telling the model to improve them did not work. I kept trying to fix the slide by re-prompting. This doom loop was quite frustrating. Once I started concentrating on improving the tooling that produced the output, I got better results. Generating the code for specific tasks like writing tests for part of the tooling or generating a specific type of data chart was fairly straightforward with today’s models (I call these “task-level” flows). Having a set of tools for agents that have sufficient flexibility but also the necessary constraints and logging all my decisions and preferences for an agent to recall later (i.e. memory) takes time and effort; and will require continued curation. I believe managing these “system-level” flows will suck up more and more time, but produce significant efficiency improvements overall for many semi-structured workflows like slide creation. The net benefit crossover point for any particular workflow, when and if it comes, is not just the tool getting better. It is also getting more structured about how I (or we) use the models.
New type of work
A couple years ago my workday consisted of three main activities: doing things, reviewing what I did, and coordinating with other people. The chart above is a rough split of how that changed. “Doing” shrank to maybe 10% of the day. Three new line items appeared: context and tooling (writing the specs, configs, and environment setup that let the model produce something useful), knowledge curation (recording decisions and preferences so the model can recall them later), and agent management (starting sessions, reading logs, restarting when the model goes in circles). Taken together, these new categories take up more than half the day. Review also grew, since verifying the model’s output takes more attention than checking your own. This shift happened because code got cheap to produce but expensive to specify well. The context documents I write for the model are often the clearest record of what a project actually does; I wrote them to steer the model, and they ended up being the thing I reach for when I come back to a project weeks later [5, 6].
Deciding what matters
The model can execute. It can write code, run tests, generate charts, and refactor modules efficiently and accurately. What it cannot do well is decide what matters: which problem to solve next, which trade-off to accept, or which output is good enough [7].
One formulation of this is that AI handles most of the work, but the remaining part is the job [9]. That description feels close to what I think is happening and will continue to happen. The cheap parts will get cheaper, but the expensive parts will become even more valuable: judgment, taste, and deciding what to write down clearly enough that someone (or something) else could do it.
References
- METR RCT: 19% slowdown for experienced open-source developers working on their own repositories, measured with early-2025 models.
- Tom Johnell: On exhaustion doom-loops in agentic workflows.
- Dan Lants: AI amplifies good engineering practices; structural investment compounds.
- Anthropic, “Building effective agents”: Start simple, add complexity only when needed.
- d11r: Theory building and understanding debt.
- Joan Westenberg: Individual accountability through documents.
- Dynomight: Taste and judgment about which factors matter.
- METR follow-up: Replication difficulties; developers refused to work without AI.
- Martynasm: Semi-decidable jobs and the 80/20 problem.
I hope to share my thoughts about healthcare data and AI, unrelated technical topics, and random things I find interesting. Do not expect any good reads!
Thanks for stopping by.
projects
Hue selectivity from recurrent circuitry
The perception of color involves a transformation from the spectral properties of visual stimuli to derived perceptual quantities such as brightness, saturation and hue. Although hue selective neurons, which respond to narrow regions in color space, have been reported in primates, they have not been identified in other species including more accessible organisms, which would facilitate circuit level analyses. Here we show that neurons in the Drosophila optic lobe have hue selective properties, with narrow tuning for both spectral and non-spectral colors. We construct a connectomics constrained circuit model that accounts for this hue selectivity. Our model, combined with genetic manipulations, shows that recurrent connections in the circuit are critical for the tuning properties of Drosophila hue selective neurons. Our findings reveal the circuit basis for a transition from physical detection to sensory perception in color vision.
Hue selectivity from recurrent circuitry, Nature Neuroscience, 2024
Presented at Columbia Neurotheory Meeting, 2022
Circuit mechanisms of chromatic encoding
Spectral information is commonly processed in the brain through generation of antagonistic responses to different wavelengths. In many species, these color opponent signals arise as early as photoreceptor terminals. Here, we measure the spectral tuning of photoreceptors in Drosophila. In addition to a previously described pathway comparing wavelengths at each point in space, we find a horizontal-cell-mediated pathway similar to that found in mammals. This pathway enables additional spectral comparisons through lateral inhibition, expanding the range of chromatic encoding in the fly. Together, these two pathways enable efficient decorrelation and dimensionality reduction of photoreceptor signals while retaining maximal chromatic information. A biologically constrained model accounts for our findings and predicts a spatio-chromatic receptive field for fly photoreceptor outputs, with a color opponent center and broadband surround. This dual mechanism combines motifs of both an insect-specific visual circuit and an evolutionarily convergent circuit architecture, endowing flies with the ability to extract chromatic information at distinct spatial resolutions.
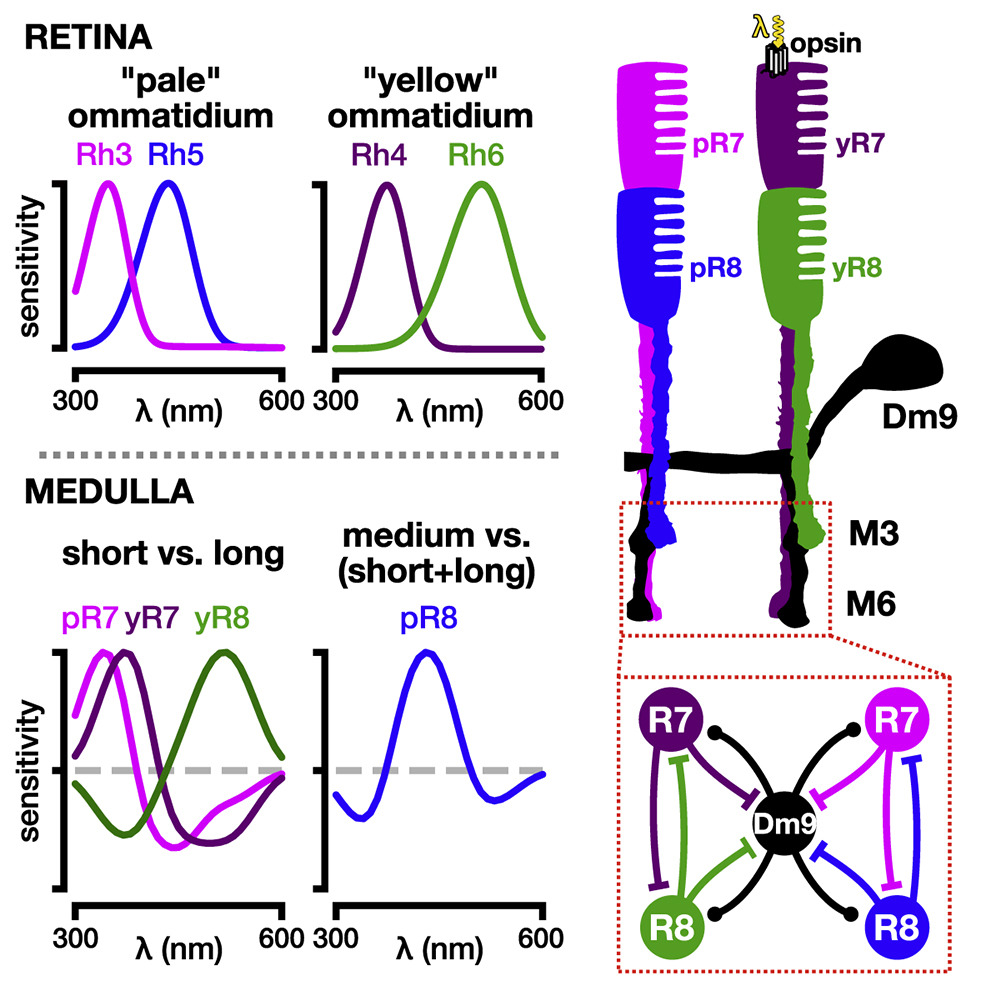
Circuit Mechanisms Underlying Chromatic Encoding in Drosophila Photoreceptors, Current Biology, 2020
Presented at Columbia Neurotheory Meeting, 2019
Normative models of spatio-spectral decorrelation
In line with the efficient coding hypothesis, the early visual system aims to minimize spectral and spatial redundancies arising from overlapping opsin sensitivities in retinal photoreceptors (PRs) and highly correlated structure in natural scenes. Encoding color information, or spectral information independent of intensity, requires comparing activities across different types of PRs. Mounting evidence shows that several species across the animal kingdom, such as the fruit fly Drosophila Melanogaster, have an uneven proportion of PR types in their retinas. However, it is unknown whether this uneven proportion is optimized for objectives relevant to the early color processing of natural scenes, as previous studies have modeled spectral and spatial processing in the early fly visual system independently. We built a collection of models incorporating both spatial and spectral information to solve tasks relevant to the fly’s early visual system, such as predictive coding at the level of PR inputs for spatial decorrelation in the retina as well as spatial and spectral decorrelation at the level of PR outputs via color opponency mechanisms. Using this framework, we asked how varying the ratio of the fly’s two main PR types changed performance accuracy on these tasks. From this normative approach, we were able to conclude that the optimal ratio of PR types to best solve these tasks aligns with the experimentally observed distribution and showed this for multiple opsin sensitivity profiles determined within and across labs. Moreover, shuffling either spatial or spectral information in the input natural scene predicted an even PR type ratio, suggesting that biologically observed PR type ratios are optimized for spectral and spatial decorrelation. Altogether, these results suggest that natural scene statistics may have shaped the ratio of PR types in the fly retina through evolutionary mechanisms, providing important implications for understanding sensory systems in an ecologically relevant context.

Normative models of spatio-spectral decorrelation predict observed receptor distributions, CoSyNe, 2022
Geometry of color spaces (dreye)
Color vision represents a vital aspect of perception that ultimately enables a wide variety of species to thrive in the natural world. However, unified methods for constructing chromatic visual stimuli in a laboratory setting are lacking. Here, we present stimulus design methods and an accompanying programming package to efficiently probe the color space of any species in which the photoreceptor spectral sensitivities are known. Our hardware-agnostic approach incorporates photoreceptor models within the framework of the principle of univariance. This enables experimenters to identify the most effective way to combine multiple light sources to create desired distributions of light, and thus easily construct relevant stimuli for mapping the color space of an organism. We include methodology to handle uncertainty of photoreceptor spectral sensitivity as well as to optimally reconstruct hyperspectral images given recent hardware advances. Our methods support broad applications in color vision science and provide a framework for uniform stimulus designs across experimental systems.
Exploiting colour space geometry for visual stimulus design across animals, Phil. Trans. R. Soc. B, 2022
Probabilistic circuit model analysis
Sensory circuits are complex systems that process information through feedforward and recurrent interactions among a set of neurons. While these circuits have been studied extensively, modeling their underlying mechanisms and computations can be challenging when not all neurons in the circuit are recorded from and when synapse information is incomplete or inaccurate. In this study, we propose a probabilistic framework that predicts the responses of unobserved neurons using anatomical constraints. We demonstrate the effectiveness of our approach on both simulated data using randomly generated connectomes with varying complexities and real neural data from the fruit fly optic lobe. Our approach accurately predicts the responses of unobserved neurons in simulated and real data. Given our probabilistic framework, we are also able to infer what types of experiments are optimal for subsequent experiments. Overall, our method provides a useful tool for modeling the complex mechanisms underlying sensory information processing in biological circuits.
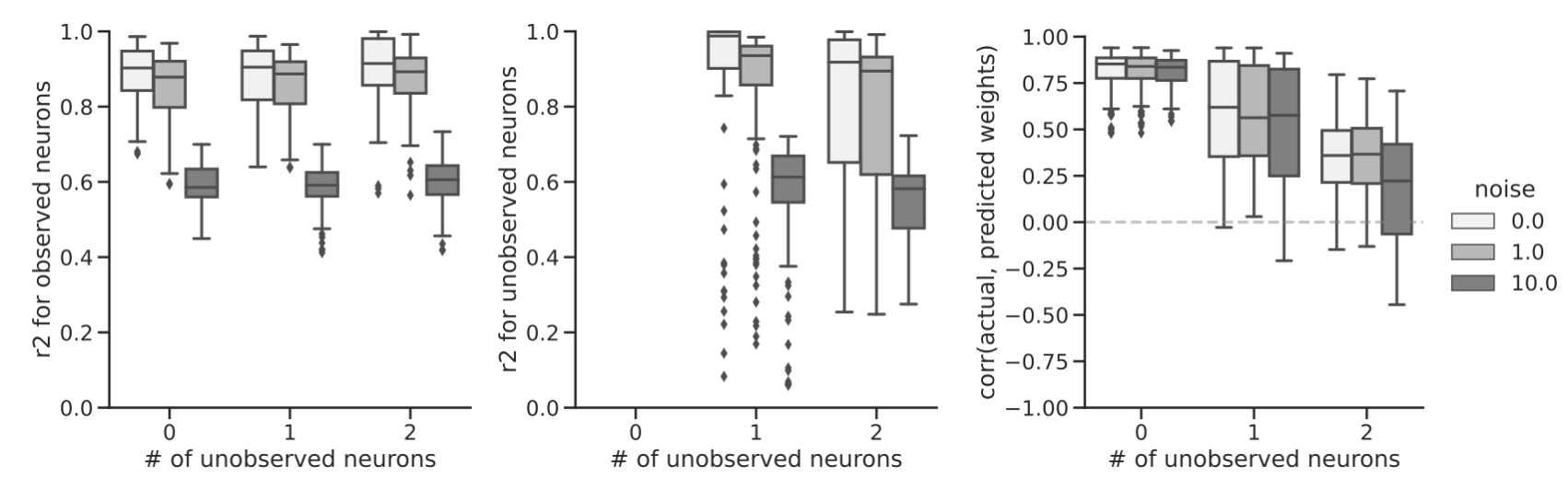
Presented at Columbia Neurotheory Meeting, 2020 and 2021
Encoding models (scidoggo)
A collection of models and data tools I built and used across my research projects. Includes constrained convex optimization procedures, interpolation models, nonlinear encoding models, and the probabilistic circuit models described above. Still under development.
Medical decisions rely on data from different sources: wearable sensors, electronic health records, and bedside monitors. Each comes with its own challenges around sequence length, irregular sampling, and individual variability. Across these projects, we are building models that learn temporal patterns from each modality, with the longer-term goal of combining them into unified patient representations.
Biometric time-series models
Conditions like infections, pneumonia, and constipation show up as multi-day patterns in wearable data, but most ML models struggle with sequences that long. At Mountain Biometrics, we worked on deep state-space model architectures that scale linearly with sequence length, learning individual “biometric profiles” to detect these healthcare events from wearable devices. The goal was to make this kind of monitoring accessible in underserved communities where it currently does not exist.
EHR foundation models
Electronic health records pose a similar challenge at a different scale: the sequences are longer, irregularly sampled, and mix different data types. We are working on a transformer-based foundation model with Mixture-of-Experts routing, trained on ~298M timeline positions from MIMIC-IV. The tokenization follows previous EHR encoding approaches: patient demographics, quantized lab values, time interval tokens, and medical event codes get mapped into a unified sequence.
Field casualty management AI
Prolonged field care combines both problems: medics need to interpret wearable biometrics over hours to days, in conditions where EHR-style clinical context is unavailable. At Mountain Biometrics, we built models for predicting individual progression of blood loss and likelihood of sepsis from heart rate, blood oxygen, and other wearable signals, designed to integrate into existing military hardware and software.
Field casualty management AI, W.W. Pettine, M. Christenson, P. Koirala, Defense TechConnect, 2023
Assessing Foundation Models’ Transferability to Physiological Signals in Precision Medicine, M. Christenson, C. Geary, B. Locke, P. Koirala, W.W. Pettine, Defense AI in Medicine, 2024
Efficient EHR Foundational Models: A Mixture-of-Experts Approach for Patient Timeline Prediction, Athreya, M. Christenson, W.W. Pettine, AI Summit, University of Utah, 2025
Agentic engineering kit
A collection of engineering standards, automated workflows, and reusable components I put together for AI-assisted software development. Includes coding principles, on-demand skills, and GitHub Actions that can be pulled in via AI assistant rules, git subtree, or individual files. Meant as a starting point that teams can adapt to how they actually work.
neuralsignal/agentic-engineering-kit
Wikipedia to YouTube shorts
In this proof of concept, I am scraping the Wikipedia article of the day to create youtube shorts from start to finish using OpenAI’s GPT model and various open-source HuggingFace models to convert text to audio, text to images, and text to video. The video encoding needs to be improved and the images aren’t nicely cut into a smooth sequence.
Movie database querying
In this proof of concept, I am trying to play around with various LLM APIs, including langchain and llamaindex, to build a semantic search over a movie database.
Auto-GPT contributions
I contributed a couple of features for this project, mainly an audio-to-text converter and image summarizer.
Obsidian export
I use Obsidian for most of my writing, so I built a tool that converts Obsidian-flavored Markdown into PDF and DOCX documents. It handles wikilinks, embeds, Mermaid diagrams, and callouts through a 5-stage processing pipeline. You can set up profiles for different styling needs.
Obsidian import
The inverse — pulls content from PDFs, Word documents, PowerPoint, spreadsheets, and other formats into Obsidian-ready Markdown with YAML frontmatter. Useful for batch-importing reference material.
Excel financial model tooling
A tool that takes a YAML spec and generates Excel workbooks with formulas, styling, and named ranges. I built it to automate the tedious parts of financial modeling — P&L statements, DCF analyses, budgets, scenario comparisons. Works as both a CLI tool and a Python API.
Loris
I built Loris as a database system for the Behnia lab at Columbia. The lab used it daily for data entry, maintenance, and analysis. It was designed for a Drosophila lab but flexible enough for other research labs. The lab has since moved to a different tech stack, so this project is no longer maintained.
Puffbird
A pandas extension for handling DataFrames with complex nested cells. I built it because I kept running into multi-valued columns that were painful to work with. Some features have become redundant since pandas added explode and similar methods, but the library still fills gaps for certain workflows. Maintained at an irregular rate.
selected works
-
+ Assessing Foundation Models’ Transferability to Physiological Signals in Precision MedicineAI in Medicine Conference, 2024
-
+ Inferring feedforward inputs to data-constrained recurrent neural networksColumbia Neurotheory Meeting, 2021
cv
- Skiing, board games, dancing, markets
about me
I lead medical data and AI efforts at Sanoptis, where I work on bringing machine learning closer to everyday clinical care. I also teach part-time as Adjunct Faculty at the University of Utah. Before that, I spent time building healthcare AI systems at MTN and studied computational neuroscience at Columbia University. Most of what I do sits at the intersection of time-series analysis, medical imaging, and making models that clinicians can actually trust and use.
When I am not working on my data and software projects, I enjoy exploring the great outdoors via gradient descent, playing board games with friends, and dancing salsa and the waltz. My long-term goal is to apply my skills in data and mathematical modeling to automate complex processes and gain a deeper understanding of how we interpret the world around us.